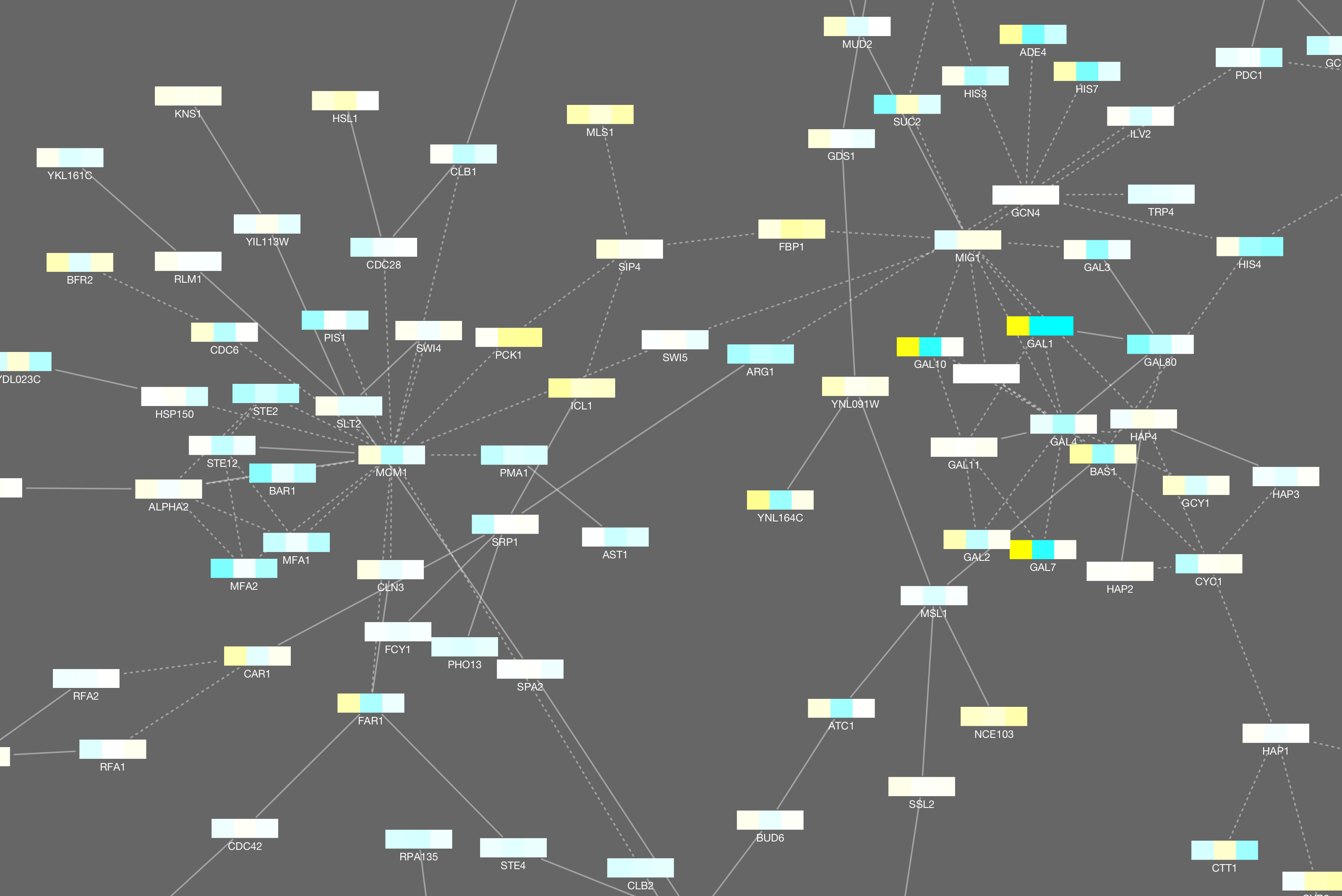
Similarity networks complement phylogenetic trees and multiple sequence alignments, two more traditional approaches generally used to study and infer information derived from comparisons of protein sequences. They have also been used to detect errors in function annotation and to study the evolution of multi-domain proteins.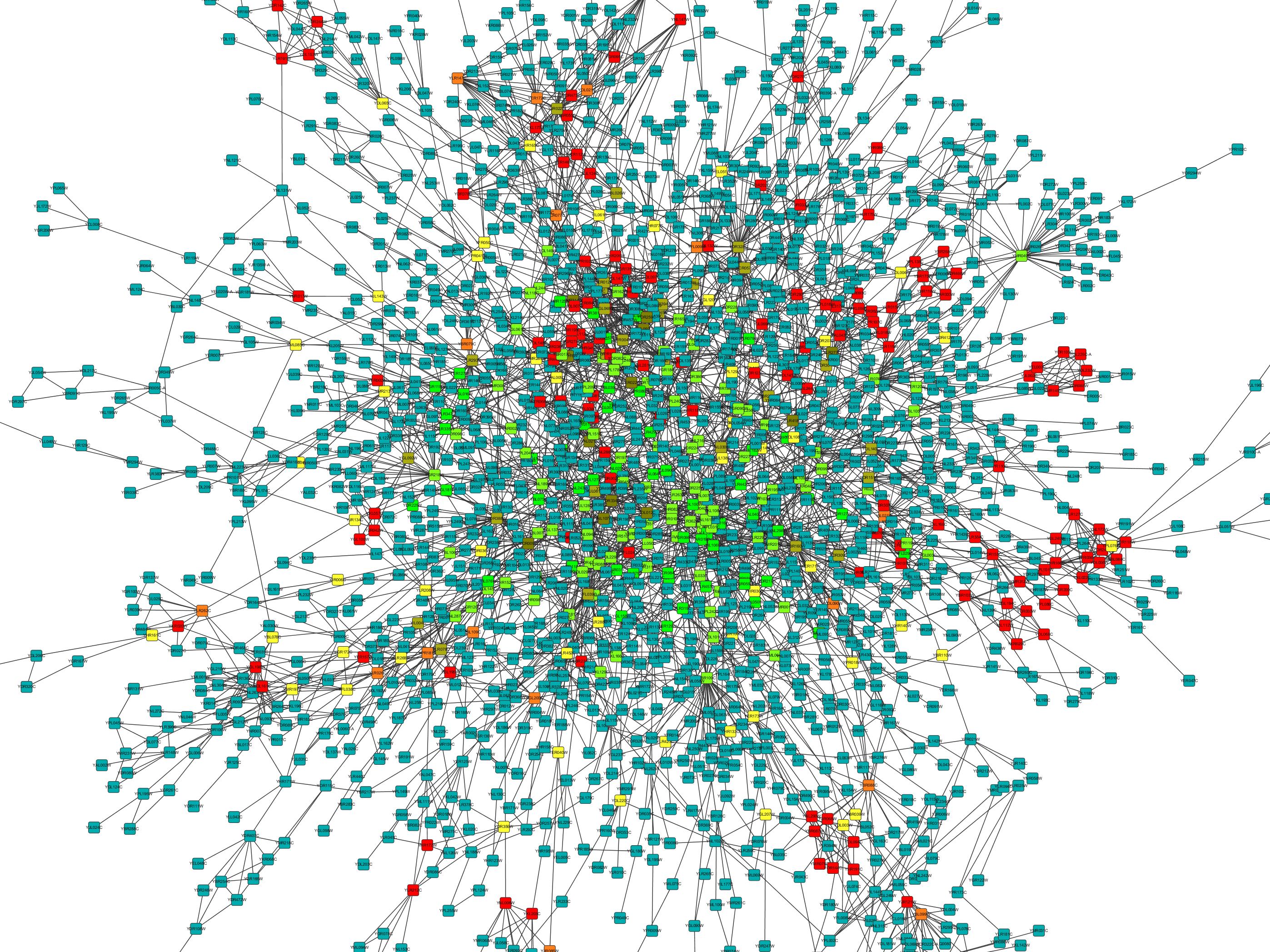 
Such networks are increasingly being used for functional and structural protein annotation. Protein sequence and structure similarity networks are bi-dimensional graphs where proteins are nodes with edges between them representing the pairwise similarity between the nodes they connect. The Phytoscape framework is available to build similarity networks, but it must be installed locally and offers a limited way to simplify large networks to be used in Cytoscape. Precomputed pairwise comparisons with functional and structural annotations are available for instance in SIMAP, but one must build a similarity network by hand from this database. Current methods still lack tools for the visualization of their results, in order to aid in their interpretation, analysis and to aid experts with curation. Most computational approaches rely on pairwise similarity to known proteins to suggest functional annotations derived by homology to annotated database entries. Recently there has also been the first Critical Assessment of Function Annotation (CAFA) experiment to assess the performance of function prediction methods. There are many ongoing projects trying to reduce the gap between known proteins and their functional annotation either computationally or experimentally. Currently, most known proteins lack any functional annotation and very little is known about the vast majority. The main protein sequence databases contain tens of millions of entries with many more sequences becoming continuously available due to the numerous genome sequencing efforts. This does not alter the authors' adherence to all the PLOS ONE policies on sharing data and materials. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.Ĭompeting interests: Co-author Silvio Tosatto is a PLOS ONE Editorial Board member. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.įunding: This project was supported by the University of Padova, FIRB Futuro in Ricerca and CARIPLO to ST. Received: JAccepted: SeptemPublished: November 12, 2013Ĭopyright: © 2013 Martin et al. PLoS ONE 8(11):Įditor: Elena Papaleo, University of Copenhagen, Denmark Citation: Martin AJM, Walsh I, Domenico TD, Mičetić I, Tosatto SCE (2013) PANADA: Protein Association Network Annotation, Determination and Analysis.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
